
SAAMBE-SEQ: A Sequence-based Method for Predicting Mutation Effect on Protein-protein Binding Affinity
Professor Emil Alexov Group



About SAAMBE-SEQ
SAAMBE-SEQ is a sequence-based machine learning algorithm to predict the binding energy changes upon single mutation in protein-protein complexes. Unlike other existing methods, SAAMBE-SEQ does not require a 3D complex structure as input. This method can be applied to protein complexes without known structure. The accuracy of SAAMBE-SEQ is comparable to existing structure-based methods. This method can be used to provide understanding the effect of mutation in protein complexes using sequence alone.
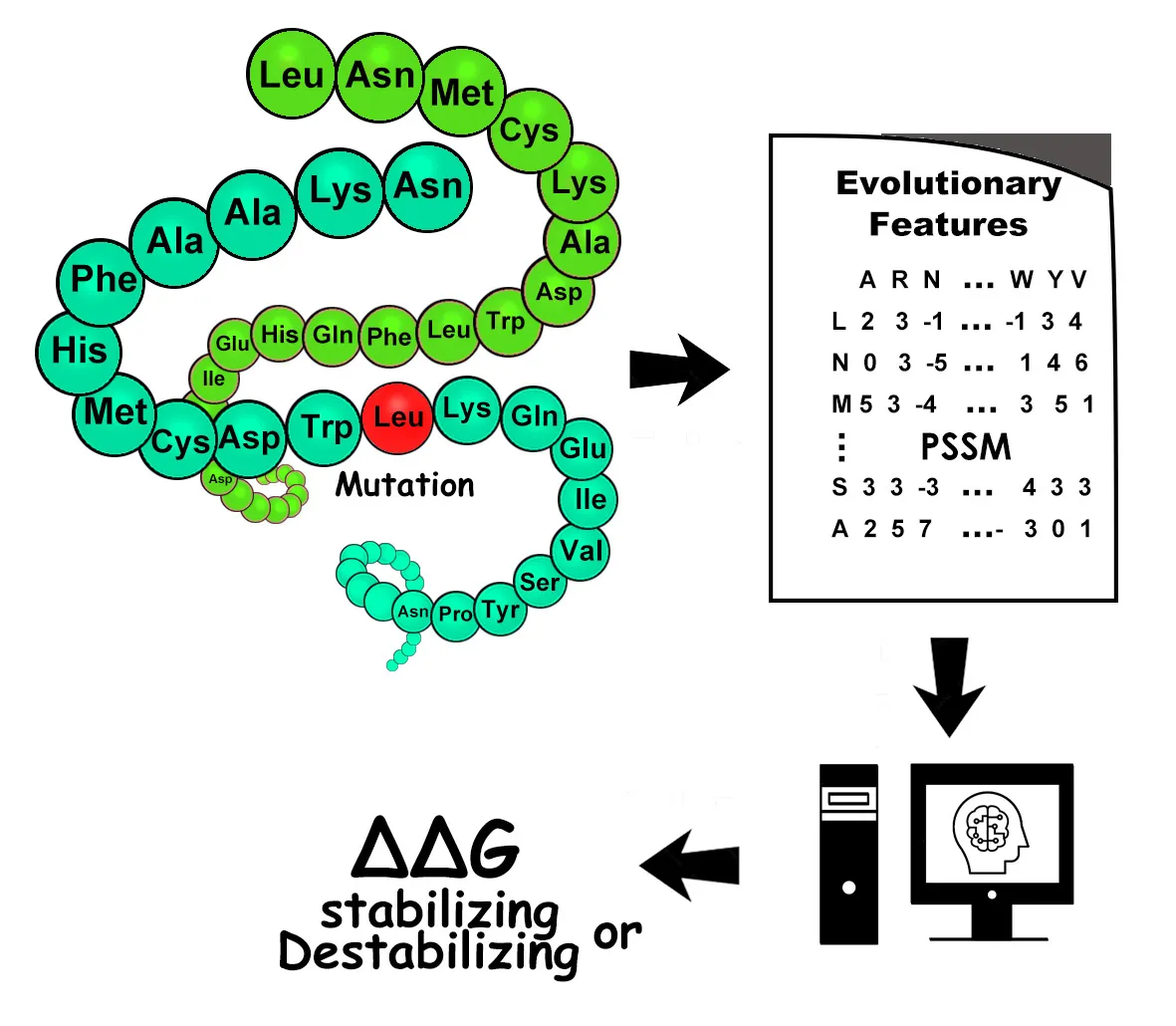
Li, G.; Pahari, S.; Murthy, A.K.; Liang, S.; Fragoza, R.; Yu, H.; Alexov, E. SAAMBE-SEQ: A Sequence-based Method for Predicting Mutation Effect on Protein-protein Binding Affinity.. Bioinformatics 2021 https://doi.org/10.1093/bioinformatics/btaa761