SAAMBE-3D: Predicting Effect of Mutations on Protein-Protein Interactions
Emil Alexov Group



About SAAMBE-3D
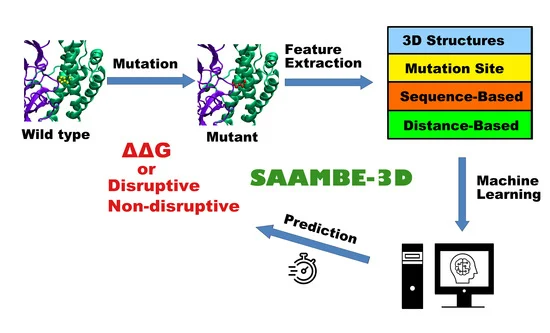
SAAMBE-3D is a newly developed machine learning algorithm to predict the effects of
single amino acid mutation on PPIs. It allows addressing two types of questions:
(a) prediction of binding free energy change caused by a mutation and
(b) prediction if mutation disrupts or not PPIs.
We also provide downloadable stand-alone code. Both codes are very fast, providing output
in a fraction of second and thus can be used for genome-scale investigations. The accuracy
of the predicting binding tree energy change was tested against SKEMPI-2 database in 5 fold test,
resulting in pearson correlation coefficient (PCC) of 0.8.
The accuracy of predicting disruptive/non-disruptive mutations was tested against
INstruct database achieving AUC=1.0