Toward understanding biological functions in a cell: Multi-scale modeling of electrostatic potential in biological systems
by Dr. Emil Alexov
 Currently, computational biophysical science addresses biological questions on various scales and details. Some researchers aim at modeling and understanding how individual molecules, proteins, DNA and RNA, perform their biological function. Such an investigation typically models biological macromolecules as objects made of many atoms connected with chemical bonds. Other researchers focus on revealing the molecular mechanism of large biological assemblies as photosynthetic machinery, respiratory complex and viruses, applying different level of details from atomistic to coarse-grained models. Furthermore, other biophysicists are interested to understand the interactions between different types of cells and tissues, and in such a case the atomistic models are not feasible.
Increasing the scale, one wants to apply biophysical methods to study the function of organs as the heart, lung, kidney and many others, which obviously cannot be done with atomistic models. The question is: how to bridge such diverse scales, since the average size of a protein is about 30 Angstroms (3 nano-meters), while the size of an organ is about 10cm and more (about 10,000,000 larger than an average molecule)?
The most promising approach to the above question is called multi-scale method such that the large object and its building blocks (molecules) are investigated with different levels of details. Our efforts in this direction are to provide more robust and efficient computer algorithms and software to allow the computational biophysics community to study biological entities of different sizes, from relatively small macromolecules to viruses.
Our current research utilizes a well-known computer package, the DelPhi package, which was originally developed in Dr. Honigs lab at Columbia University in New York. The efforts in the lab are directed to modernize Delphi and to make it capable of calculating electrostatic potential and energies of large systems as viruses and mitochondrial complex. This is done by developing parallelized schemes that dramatically reduce the computational time and computer memory requirements.
The uniqueness of our approach is that it employs different techniques depending on the physical nature of the problem, which needs to be parallelized. Thus, we have developed new multi-level (parallelization is achieved at different levels of the algorithm) and interleaved (parallelization is implemented by interleaving the computational tasks) method, the MLIPB method, to parallelize standard methods for computing electrostatics potential and energies in the framework of the Poisson-Boltzmann equation.
The leading scientist in this project, Dr. Chuan Li, a postdoctoral researcher at the Computational Biophysics Laboratory in the Department of Physics and Astronomy, revisited the existing DelPhi algorithm to group the algorithmic tasks into several distinctive classes depending on the physical process being modeled. This was a very important development because he managed to increase the computational speed of calculations without losing accuracy.
In other words, the parallelized DelPhi is almost 100 times faster than the serial one, while retaining the same accuracy. From point of view of biological applications, this translates into a tool that can mole small and large objects with the same accuracy, allowing viruses and large macromolecular assemblages to be investigated at the atomic level of detail.
The ability of precise modeling of electrostatic potential, energy and forces is crucial for understanding biological processes taking place on various scales. The reason for that is that each atom within a protein, a DNA or a RNA carries electrostatic charge and at the same time the atoms are positioned within very short distances among themselves.
Applying high-school physics formulas, the Coulomb Law of Electrostatics, reveals that the strength of interactions at such distances is greater than any other force. Because of that, our development provides the Biophysics Community with a tool that enables revealing the role of electrostatics in very large systems bridging the gap between understanding the function of individual biological macromolecules and the function of much larger entities made of many proteins, DNAs and RNAs.
Our research is highly relevant to any study involving biological macromolecules, their assemblages and even larger entities. It paves the way of understanding the molecular mechanism associated with various biological functions in the cell and in a long run the role of electrostatics in process occurring in various organs. This is also relevant to revealing the effects of genetic disorders and the corresponding mechanism of the diseases, providing the opportunity the disease effect to be modeled at various scales and details.
Our goal in a long run is to further increase our capability to model even larger objects as components of the cell and the entire cell on level of details which will allow for meaningful biological conclusions to be made. This will be done by gluing individual atoms into larger objects, called groups of atoms, which then will make the structure of the large object (for example cellular compartments or the entire cell).
Currently the lab is progressing very well in this direction, the leading researcher being Mr. Nicholas Smith, a graduate student in the lab, who developed a tool called Protein-Nano Object Integrator. This tool allows for generating 3D-objects as multitude of pseudo-atoms and manipulating the objects in real time.
Currently, computational biophysical science addresses biological questions on various scales and details. Some researchers aim at modeling and understanding how individual molecules, proteins, DNA and RNA, perform their biological function. Such an investigation typically models biological macromolecules as objects made of many atoms connected with chemical bonds. Other researchers focus on revealing the molecular mechanism of large biological assemblies as photosynthetic machinery, respiratory complex and viruses, applying different level of details from atomistic to coarse-grained models. Furthermore, other biophysicists are interested to understand the interactions between different types of cells and tissues, and in such a case the atomistic models are not feasible.
Increasing the scale, one wants to apply biophysical methods to study the function of organs as the heart, lung, kidney and many others, which obviously cannot be done with atomistic models. The question is: how to bridge such diverse scales, since the average size of a protein is about 30 Angstroms (3 nano-meters), while the size of an organ is about 10cm and more (about 10,000,000 larger than an average molecule)?
The most promising approach to the above question is called multi-scale method such that the large object and its building blocks (molecules) are investigated with different levels of details. Our efforts in this direction are to provide more robust and efficient computer algorithms and software to allow the computational biophysics community to study biological entities of different sizes, from relatively small macromolecules to viruses.
Our current research utilizes a well-known computer package, the DelPhi package, which was originally developed in Dr. Honigs lab at Columbia University in New York. The efforts in the lab are directed to modernize Delphi and to make it capable of calculating electrostatic potential and energies of large systems as viruses and mitochondrial complex. This is done by developing parallelized schemes that dramatically reduce the computational time and computer memory requirements.
The uniqueness of our approach is that it employs different techniques depending on the physical nature of the problem, which needs to be parallelized. Thus, we have developed new multi-level (parallelization is achieved at different levels of the algorithm) and interleaved (parallelization is implemented by interleaving the computational tasks) method, the MLIPB method, to parallelize standard methods for computing electrostatics potential and energies in the framework of the Poisson-Boltzmann equation.
The leading scientist in this project, Dr. Chuan Li, a postdoctoral researcher at the Computational Biophysics Laboratory in the Department of Physics and Astronomy, revisited the existing DelPhi algorithm to group the algorithmic tasks into several distinctive classes depending on the physical process being modeled. This was a very important development because he managed to increase the computational speed of calculations without losing accuracy.
In other words, the parallelized DelPhi is almost 100 times faster than the serial one, while retaining the same accuracy. From point of view of biological applications, this translates into a tool that can mole small and large objects with the same accuracy, allowing viruses and large macromolecular assemblages to be investigated at the atomic level of detail.
The ability of precise modeling of electrostatic potential, energy and forces is crucial for understanding biological processes taking place on various scales. The reason for that is that each atom within a protein, a DNA or a RNA carries electrostatic charge and at the same time the atoms are positioned within very short distances among themselves.
Applying high-school physics formulas, the Coulomb Law of Electrostatics, reveals that the strength of interactions at such distances is greater than any other force. Because of that, our development provides the Biophysics Community with a tool that enables revealing the role of electrostatics in very large systems bridging the gap between understanding the function of individual biological macromolecules and the function of much larger entities made of many proteins, DNAs and RNAs.
Our research is highly relevant to any study involving biological macromolecules, their assemblages and even larger entities. It paves the way of understanding the molecular mechanism associated with various biological functions in the cell and in a long run the role of electrostatics in process occurring in various organs. This is also relevant to revealing the effects of genetic disorders and the corresponding mechanism of the diseases, providing the opportunity the disease effect to be modeled at various scales and details.
Our goal in a long run is to further increase our capability to model even larger objects as components of the cell and the entire cell on level of details which will allow for meaningful biological conclusions to be made. This will be done by gluing individual atoms into larger objects, called groups of atoms, which then will make the structure of the large object (for example cellular compartments or the entire cell).
Currently the lab is progressing very well in this direction, the leading researcher being Mr. Nicholas Smith, a graduate student in the lab, who developed a tool called Protein-Nano Object Integrator. This tool allows for generating 3D-objects as multitude of pseudo-atoms and manipulating the objects in real time.
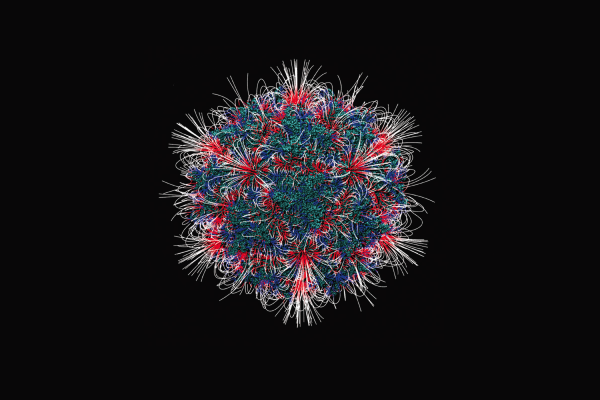
Image Caption: the image shows the distribution of electrostatic field mapped onto the surface of the adeno-associated virus, PDB ID 3KIC. For media inquiries, please contact Brian Mullen at mullen2@clemson.edu or 864-656-2063.