SAMPDI-3D: Predicting protein-DNA binding free energy change upon mutations
Emil Alexov Group



About SAMPDI-3D
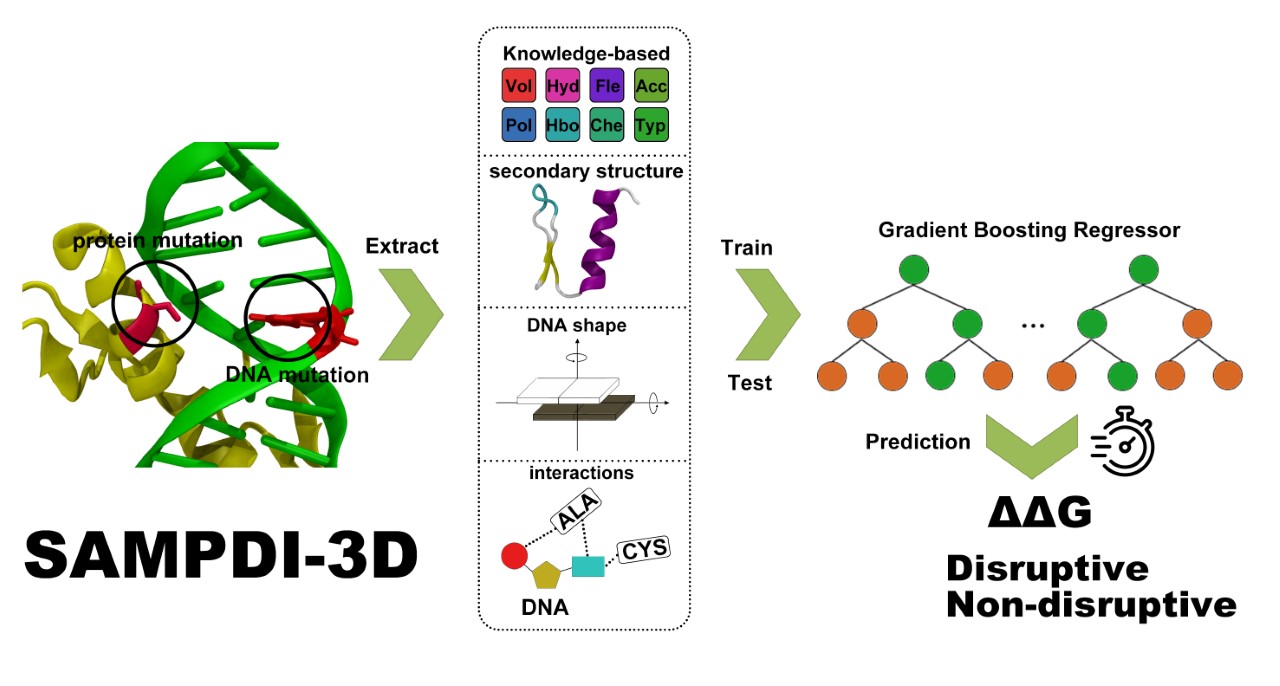
SAMPDI-3D uses a gradient boosting decision tree machine learning algorithm with features as physicochemical properties, structure of mutation site and protein-DNA interactions to predict the change of binding free energy. SAMPDI-3D outperforms all existing state-of-the-art methods in both the Pearson correlation coefficient and root-mean-squared-error parameters for several independent datasets(available here). Users can also download SAMPDI-3D stand alone code (available here) and use it locally.
Cite "Bioinformatics, 2021 Aug 3;btab567. doi: 10.1093/bioinformatics/btab567"