
SAAFEC-SEQ: an online application for calculating folding free energy changes in proteins caused by missense mutations
Professor Emil Alexov Group



About SAAFEC-SEQ
SAAFEC-SEQ uses the PsePSSM algorithm to predict the stability changes upon a single mutation in protein. SAAFEC-SEQ uses gradient boosting decision tree repressor to encode physicochemical properties, sequence features and evolutionary information to compute the change in stability free energy resulting from single mutations. SAAFEC-SEQ outperforms all existing state-of-the-art sequence-based methods in both the Pearson correlation coefficient and root-mean-squared-error parameters for several independent datasets(available here). SAAFEC-SEQ (available here) and use it locally.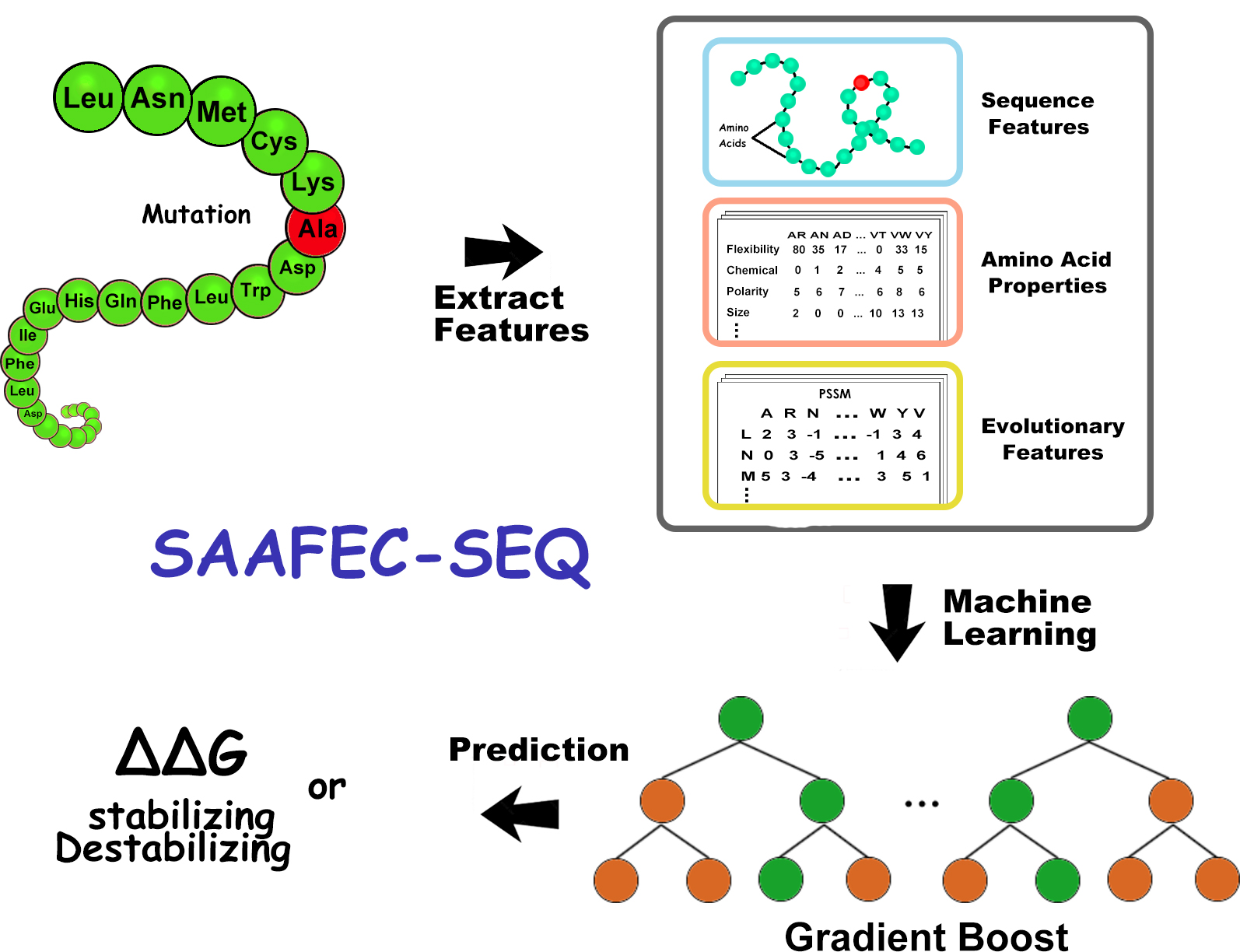
Li, G.; Panday, S.K.; Alexov, E. SAAFEC-SEQ: A Sequence-Based Method for Predicting the Effect of Single Point Mutations on Protein Thermodynamic Stability. Int. J. Mol. Sci. 2021, 22, 606. https://doi.org/10.3390/ijms22020606
uniref100 database, which is a clustered version of nr database to make computations less intensive. Note that results may not be identical to that for nr database. However, you could download standalone version (available here) and use nr database on your own computer to achieve better accuracy.